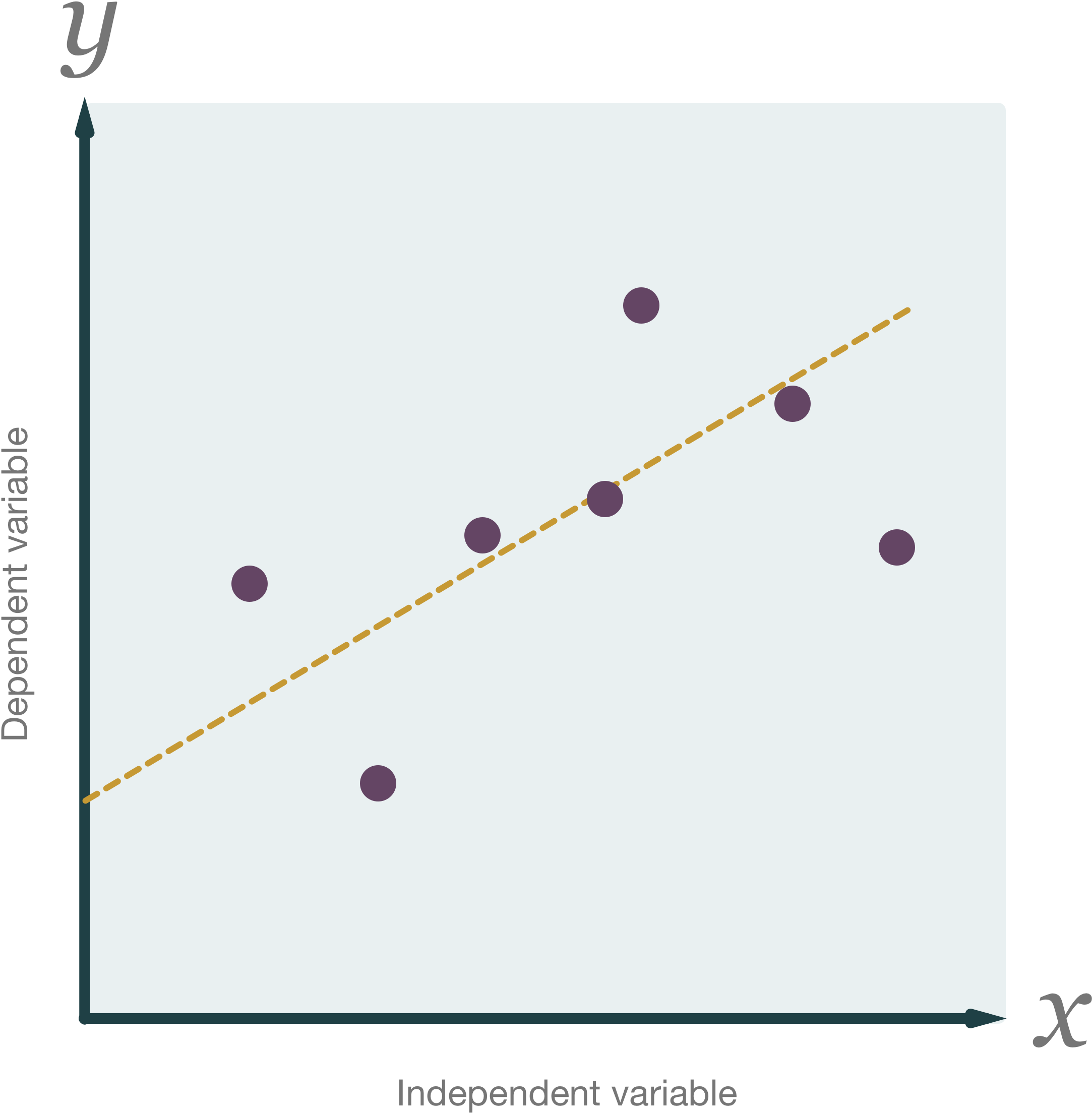
Overview
In statistics, linear regression is an approach for modelling the relationship between a scalar dependent variable y and one or more explanatory variables (or independent variables) denoted X. In linear regression, the relationships are modeled using linear predictor functions whose unknown model parameters are estimated from the data. Such models are called linear models.
Implementation of Harp-DAAL-LinReg
Harp-DAAL-LineReg is built upon the original DAAL distributed implementation of the algorithm using the Harp interface libraries for the collective-communication among the data nodes. The implementation of LinReg in our Harp-DAAL consists of two levels. At the top Level, using Harp’s reduce operation to communication model data among mappers. At the bottom Level, using DAAL’s kernels to conduct local computations.
Brief background
- There are primarily three kinds of nodes involved with the implementation.
- name node
- data node
- master node
- slave node
The Java language services provided by Intel as a wrapper to their C++ code.
The description of the intel daal’s JAVA API used can be found here
Code Walk-Through
The actual implemented code can be found here. The MapCollective function is defined here.
Step 1 (on data nodes)
The first step involves reading the files from the hdfs filesystem after splitting files between each mapper. Splitting is done by MultipleFileInputFormat class defined here. Data is converted into array which is eventually converted into the daal Numeric Table datastructure. In this example the files have been stored on the hdfs. The files have to be read in the daal table format as the local computations are performed by the daal libraries. Getting array from csv file -
public static List<double[]> loadPoints(String file,
int pointsPerFile, int cenVecSize,int labelSize,
Configuration conf) throws Exception {
System.out.println("filename: "+file );
List<double[]> points = new LinkedList<double[]>();
double[] trainingData =
new double[pointsPerFile * cenVecSize];
double[] labelData =
new double[pointsPerFile * labelSize];
Path pointFilePath = new Path(file);
FileSystem fs =
pointFilePath.getFileSystem(conf);
FSDataInputStream in = fs.open(pointFilePath);
int k =0;
int p = 0;
try{
for(int i = 0; i < pointsPerFile;i++){
String[] line = in.readLine().split(",");
for(int j = 0; j < cenVecSize; j++){
trainingData[k] = Double.parseDouble(line[j]);
k++;
}
for(int s = cenVecSize; s < cenVecSize+labelSize; s++){
labelData[p] = Double.parseDouble(line[s]);
p++;
}
}
} finally{
in.close();
points.add(trainingData);
points.add(labelData);
}
return points;
}
Converting array to Numeric Table -
List<List<double[]>> pointArrays = LinRegUtil.loadPoints(trainingDataFiles, pointsPerFile,
vectorSize, nDependentVariables, conf, numThreads);
List<double[]> featurePoints = new LinkedList<>();
for(int i = 0; i<pointArrays.size(); i++){
featurePoints.add(pointArrays.get(i).get(0));
}
List<double[]> labelPoints = new LinkedList<>();
for(int i = 0; i<pointArrays.size(); i++){
labelPoints.add(pointArrays.get(i).get(1));
}
for(int k=0;k<featurePoints.size();k++)
{
array_data_feature[k] = featurePoints.get(k);
array_startP_feature[k] = totalLengthFeature;
totalLengthFeature += featurePoints.get(k).length;
}
for(int k=0;k<labelPoints.size();k++)
{
array_data_label[k] = labelPoints.get(k);
array_startP_label[k] = totalLengthLabel;
totalLengthLabel += labelPoints.get(k).length;
}
long featuretableSize = totalLengthFeature/nFeature;
long labeltableSize = totalLengthLabel/nLabel;
//initializing Numeric Table
NumericTable featureArray_daal = new HomogenNumericTable(daal_Context, Double.class, nFeature, featuretableSize, NumericTable.AllocationFlag.DoAllocate);
NumericTable labelArray_daal = new HomogenNumericTable(daal_Context, Double.class, nLabel, labeltableSize, NumericTable.AllocationFlag.DoAllocate);
Step 2 (on data nodes)
TrainingDistributedStep1Local is created for local computation. Computed partial results are sent to master node by reduce function.
TrainingDistributedStep1Local linearRegressionTraining = new TrainingDistributedStep1Local(daal_Context, Float.class,
TrainingMethod.qrDense);
linearRegressionTraining.input.set(TrainingInputId.data, trainData);
linearRegressionTraining.input.set(TrainingInputId.dependentVariable, trainDependentVariables);
PartialResult pres = linearRegressionTraining.compute();
ts2 = System.currentTimeMillis();
compute_time += (ts2 - ts1);
ts1 = System.currentTimeMillis();
partialResultTable.addPartition(new Partition<>(this.getSelfID(), serializePartialResult(pres)));
boolean reduceStatus = false;
reduceStatus = this.reduce("linreg", "sync-partialresult", partialResultTable, this.getMasterID());
Step 3 (only on master nodes)
Final training result and model are computed on master node.
if(this.isMaster()){
TrainingDistributedStep2Master linearRegressionTrainingMaster = new TrainingDistributedStep2Master(daal_Context, Float.class,
TrainingMethod.qrDense);
ts1 = System.currentTimeMillis();
int[] pid = partialResultTable.getPartitionIDs().toIntArray();
for(int j = 0; j< pid.length; j++){
try {
linearRegressionTrainingMaster.input.add(MasterInputId.partialModels,
deserializePartialResult(partialResultTable.getPartition(pid[j]).get()));
} catch (Exception e)
{
System.out.println("Fail to deserilize partialResultTable" + e.toString());
e.printStackTrace();
}
}
ts2 = System.currentTimeMillis();
comm_time += (ts2 - ts1);
ts1 = System.currentTimeMillis();
linearRegressionTrainingMaster.compute();
trainingResult = linearRegressionTrainingMaster.finalizeCompute();
ts2 = System.currentTimeMillis();
compute_time += (ts2 - ts1);
model = trainingResult.get(TrainingResultId.model);
}
Step 4 (on master node)
PredictionResult is computed on master node.
PredictionBatch linearRegressionPredict = new PredictionBatch(daal_Context, Float.class, PredictionMethod.defaultDense);
NumericTable testData = getNumericTableHDFS(daal_Context, conf, testFilePath, 10, 250);
linearRegressionPredict.input.set(PredictionInputId.data, testData);
linearRegressionPredict.input.set(PredictionInputId.model, model);
/* Compute the prediction results */
ts1 = System.currentTimeMillis();
predictionResult = linearRegressionPredict.compute();
results = predictionResult.get(PredictionResultId.prediction);
Running the codes
Make sure that the code is placed in the /harp/ml/daal directory.
Run the harp-daal-linreg.sh script here to run the code.
cd $HARP_ROOT/ml/daal
./test_scripts/harp-daal-linreg.sh
The details of script is here