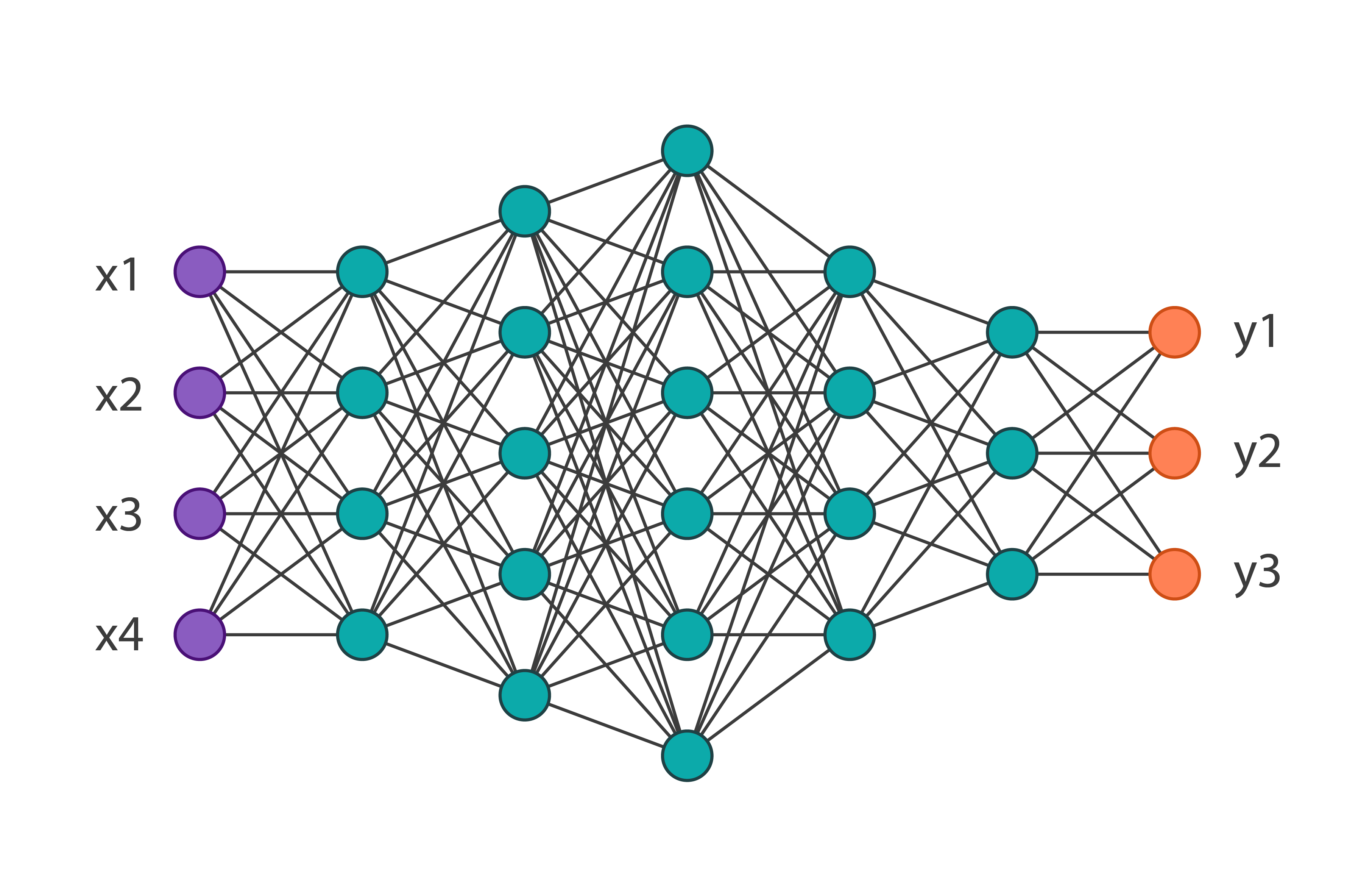
Neural Networks are a beautiful biologically-inspired programming paradigm which enable a computer to learn from observational data. The motivation for the development of neural network technology stemmed from the desire to develop an artificial system that could perform “intelligent” tasks similar to those performed by the human brain. Neural networks, with their remarkable ability to derive meaning from complicated or imprecise data, can be used to extract patterns and detect trends that are too complex to be noticed by either humans or other computer techniques.
Harp-DAAL currently supports distributed mode of Neural Networks for dense input datasets.
More algorithmic details from Intel DAAL documentation is here.